David H. Koch Professor of Science
Professor of Biology and Biological Engineering
Director, MIT Center for Precision Cancer Medicine
Director, KI Clinical Investigator Program
Contact Information
Research Areas
Immunology & immunotherapy, Nano-based drugs, Precision medicine
I am fascinated with how protein kinase signaling pathways activated by DNA damage, cell injury, and inflammation within tumors and their surrounding microenvironment control the behavior of cancers and their response to treatment.
Research Summary
Cancer cells respond to chemotherapies by activating specific signaling pathways that control whether damaged cells survive or die, and whether injured cells enhance or suppress anti-tumor immune response. Using various benchtop and computational/bioinformatic approaches, we study how signaling pathways that control DNA damage responses, survival, cell cycle progression and immune activation are integrated at the molecular and systems levels to control tumor outcomes after treatment. Clinically, our work translates into 'systems pharmacology,' namely, determining how to target cross-talk between pathways with specific drug combinations to increase cancer treatment efficacy.
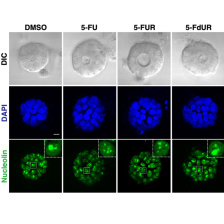
Current interests include understanding how: 1) the p38MAPK-MAPKAP Kinase-2 pathway regulates DNA damage responses in p53-mutant tumors, including its roles in cytokine signaling, drug resistance and anti-tumor immunity in ovarian cancer; 2) signaling from R-loops induced by Brd4 inhibition or RNA-binding protein disruption can be used to kill tumor cells defective in specific DNA repair pathways; 3) specific DNA repair pathways are selected after a double-strand DNA break, and whether these pathways can be targeted to improve chemotherapy responses; 4) DNA damage can induce an 'immunogenic cell injury' response that enhances anti-cancer immunotherapy; and 5) drugs such as Polo-like kinase-1 inhibitors or 5-FU can be used to treat tumors by interfering with mitotic progression or inducing an RNA damage response.
Biography
Michael B. Yaffe is the David H. Koch Professor of Science and Professor of Biology and Biological Engineering at MIT, where he has been a member of the faculty since 2000. He is also an attending surgeon and intensivist at the Beth Israel Deaconess Medical Center where he specializes in injury and surgical oncology. He has been the Director of the MIT Center for Precision Cancer Medicine since its inception. Yaffe earned his MD-PhD degree from Case Western Reserve University. He then completed a residency in General Surgery at University Hospitals of Cleveland and New England Deaconess Hospital, and a fellowship in Surgical Critical Care, Trauma and Burns at Harvard Medical School's Harvard-Longwood Critical Care Program. He was a post-doctoral fellow with Lewis Cantley in the Department of Cell Biology at Harvard Medical School where he was supported by a Physician-Scientist Fellowship from the HHMI, and a Physician-Scientist Award from the Burroughs-Wellcome Fund. Yaffe is the Scientific Editor-in-Chief of Science Signaling and a member of the Editorial Boards of Molecular & Cellular Proteomics and Cell Cycle. In 2021, Professor Yaffe was elected to the Association of American Physicians and was named a MacVicar Faculty Fellow. He is a decorated veteran of the war in Afghanistan and retired as a Colonel from the U.S. Army Reserve Medical Corps in 2021. Yaffe is a co-founder of Consensus Pharmaceuticals, the DNA Repair Company, On-Q-ity, and Merrimack Pharmaceuticals. He is also a co-founder and a member of the scientific advisory board of Applied Biomath and Thrombo-Therapeutics, and a clinical advisor to Cardiff Oncology.